Prerequisites
Outline
- Section 1: Dataset Preparation
- Section 2: Linear SVM Formulation
- Section 3: SVM Optimization using SGD
- Section 4: Model Evaluation via T-fold Cross Validation
Support Vector Machines (SVMs)
In this notebook, we will learn a linear and kernalised method of SVMs, which can be used for both regression and classification. To start with, we will focus on binary classification. We will use stochastic gradient descent (SGD) for the optimisation of the hinge loss.
We will work with the Breast Cancer Wisconsin (Diagnostic) Data Set, which you first need to download and then load in this notebook. If you faced difficulties downloading this data set from Kaggle, you should download the file directly from Blackboard. The data set contains various aspects of cell nuclei of breast screening images of patients with (malignant) and without (benign) breast cancer. Our goal is to build a classification model that can take these aspects of an unseen breast screening image, and classify it as either malignant or benign.
Section 1: Data Preparation
If you run this notebook locally on your machine, you will simply need to place the csv file in the same directory as this notebook.
If you run this notebook on Google Colab, you will need to use
from google.colab import files
upload = files.upload()
and then upload it from your local downloads directory.
# necessary imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#from google.colab import files
#upload = files.upload()
data = pd.read_csv('./data.csv')
# print shape and last 10 rows
print(data.shape)
data.tail(10)
(569, 33)
| id | diagnosis | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | ... | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | Unnamed: 32 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 559 | 925291 | B | 11.51 | 23.93 | 74.52 | 403.5 | 0.09261 | 0.10210 | 0.11120 | 0.04105 | ... | 37.16 | 82.28 | 474.2 | 0.12980 | 0.25170 | 0.3630 | 0.09653 | 0.2112 | 0.08732 | NaN |
| 560 | 925292 | B | 14.05 | 27.15 | 91.38 | 600.4 | 0.09929 | 0.11260 | 0.04462 | 0.04304 | ... | 33.17 | 100.20 | 706.7 | 0.12410 | 0.22640 | 0.1326 | 0.10480 | 0.2250 | 0.08321 | NaN |
| 561 | 925311 | B | 11.20 | 29.37 | 70.67 | 386.0 | 0.07449 | 0.03558 | 0.00000 | 0.00000 | ... | 38.30 | 75.19 | 439.6 | 0.09267 | 0.05494 | 0.0000 | 0.00000 | 0.1566 | 0.05905 | NaN |
| 562 | 925622 | M | 15.22 | 30.62 | 103.40 | 716.9 | 0.10480 | 0.20870 | 0.25500 | 0.09429 | ... | 42.79 | 128.70 | 915.0 | 0.14170 | 0.79170 | 1.1700 | 0.23560 | 0.4089 | 0.14090 | NaN |
| 563 | 926125 | M | 20.92 | 25.09 | 143.00 | 1347.0 | 0.10990 | 0.22360 | 0.31740 | 0.14740 | ... | 29.41 | 179.10 | 1819.0 | 0.14070 | 0.41860 | 0.6599 | 0.25420 | 0.2929 | 0.09873 | NaN |
| 564 | 926424 | M | 21.56 | 22.39 | 142.00 | 1479.0 | 0.11100 | 0.11590 | 0.24390 | 0.13890 | ... | 26.40 | 166.10 | 2027.0 | 0.14100 | 0.21130 | 0.4107 | 0.22160 | 0.2060 | 0.07115 | NaN |
| 565 | 926682 | M | 20.13 | 28.25 | 131.20 | 1261.0 | 0.09780 | 0.10340 | 0.14400 | 0.09791 | ... | 38.25 | 155.00 | 1731.0 | 0.11660 | 0.19220 | 0.3215 | 0.16280 | 0.2572 | 0.06637 | NaN |
| 566 | 926954 | M | 16.60 | 28.08 | 108.30 | 858.1 | 0.08455 | 0.10230 | 0.09251 | 0.05302 | ... | 34.12 | 126.70 | 1124.0 | 0.11390 | 0.30940 | 0.3403 | 0.14180 | 0.2218 | 0.07820 | NaN |
| 567 | 927241 | M | 20.60 | 29.33 | 140.10 | 1265.0 | 0.11780 | 0.27700 | 0.35140 | 0.15200 | ... | 39.42 | 184.60 | 1821.0 | 0.16500 | 0.86810 | 0.9387 | 0.26500 | 0.4087 | 0.12400 | NaN |
| 568 | 92751 | B | 7.76 | 24.54 | 47.92 | 181.0 | 0.05263 | 0.04362 | 0.00000 | 0.00000 | ... | 30.37 | 59.16 | 268.6 | 0.08996 | 0.06444 | 0.0000 | 0.00000 | 0.2871 | 0.07039 | NaN |
10 rows × 33 columns
We can see that our data set has 569 samples and 33 columns. The column id can be taken as an index for our pandas dataframe and diagnosis is the label (either M: malignant or B: benign).
Let’s prepare the data set first of all by (i) cleaning it, (ii) separating label from features, and (iii) splitting it into train and test sets.
# drop last column (extra column added by pd)
data_1 = data.drop(data.columns[-1], axis=1)
# set column id as dataframe index
data_2 = data_1.set_index(data['id']).drop(data_1.columns[0], axis=1)
# check
data_2.tail()
| diagnosis | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | ... | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||||
| 926424 | M | 21.56 | 22.39 | 142.00 | 1479.0 | 0.11100 | 0.11590 | 0.24390 | 0.13890 | 0.1726 | ... | 25.450 | 26.40 | 166.10 | 2027.0 | 0.14100 | 0.21130 | 0.4107 | 0.2216 | 0.2060 | 0.07115 |
| 926682 | M | 20.13 | 28.25 | 131.20 | 1261.0 | 0.09780 | 0.10340 | 0.14400 | 0.09791 | 0.1752 | ... | 23.690 | 38.25 | 155.00 | 1731.0 | 0.11660 | 0.19220 | 0.3215 | 0.1628 | 0.2572 | 0.06637 |
| 926954 | M | 16.60 | 28.08 | 108.30 | 858.1 | 0.08455 | 0.10230 | 0.09251 | 0.05302 | 0.1590 | ... | 18.980 | 34.12 | 126.70 | 1124.0 | 0.11390 | 0.30940 | 0.3403 | 0.1418 | 0.2218 | 0.07820 |
| 927241 | M | 20.60 | 29.33 | 140.10 | 1265.0 | 0.11780 | 0.27700 | 0.35140 | 0.15200 | 0.2397 | ... | 25.740 | 39.42 | 184.60 | 1821.0 | 0.16500 | 0.86810 | 0.9387 | 0.2650 | 0.4087 | 0.12400 |
| 92751 | B | 7.76 | 24.54 | 47.92 | 181.0 | 0.05263 | 0.04362 | 0.00000 | 0.00000 | 0.1587 | ... | 9.456 | 30.37 | 59.16 | 268.6 | 0.08996 | 0.06444 | 0.0000 | 0.0000 | 0.2871 | 0.07039 |
5 rows × 31 columns
We do a bit more preparation by converting the categorical labels into 1 for M and -1 for B.
# convert categorical labels to numbers
diag_map = {'M': 1.0, 'B': -1.0}
data_2['diagnosis'] = data_2['diagnosis'].map(diag_map)
# put labels and features in different dataframes
y = data_2.loc[:, 'diagnosis'] # loc is typically used for label indexing
X = data_2.iloc[:, 1:] # iloc is used for integer indexing
# check
print(y.tail())
X.tail()
id
926424 1.0
926682 1.0
926954 1.0
927241 1.0
92751 -1.0
Name: diagnosis, dtype: float64
| radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | fractal_dimension_mean | ... | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||||
| 926424 | 21.56 | 22.39 | 142.00 | 1479.0 | 0.11100 | 0.11590 | 0.24390 | 0.13890 | 0.1726 | 0.05623 | ... | 25.450 | 26.40 | 166.10 | 2027.0 | 0.14100 | 0.21130 | 0.4107 | 0.2216 | 0.2060 | 0.07115 |
| 926682 | 20.13 | 28.25 | 131.20 | 1261.0 | 0.09780 | 0.10340 | 0.14400 | 0.09791 | 0.1752 | 0.05533 | ... | 23.690 | 38.25 | 155.00 | 1731.0 | 0.11660 | 0.19220 | 0.3215 | 0.1628 | 0.2572 | 0.06637 |
| 926954 | 16.60 | 28.08 | 108.30 | 858.1 | 0.08455 | 0.10230 | 0.09251 | 0.05302 | 0.1590 | 0.05648 | ... | 18.980 | 34.12 | 126.70 | 1124.0 | 0.11390 | 0.30940 | 0.3403 | 0.1418 | 0.2218 | 0.07820 |
| 927241 | 20.60 | 29.33 | 140.10 | 1265.0 | 0.11780 | 0.27700 | 0.35140 | 0.15200 | 0.2397 | 0.07016 | ... | 25.740 | 39.42 | 184.60 | 1821.0 | 0.16500 | 0.86810 | 0.9387 | 0.2650 | 0.4087 | 0.12400 |
| 92751 | 7.76 | 24.54 | 47.92 | 181.0 | 0.05263 | 0.04362 | 0.00000 | 0.00000 | 0.1587 | 0.05884 | ... | 9.456 | 30.37 | 59.16 | 268.6 | 0.08996 | 0.06444 | 0.0000 | 0.0000 | 0.2871 | 0.07039 |
5 rows × 30 columns
As with any data set that has features over different ranges, it’s required to standardise the data before.
## EDIT THIS FUNCTION
def standardise(X):
mu = np.mean(X, 0)
sigma = np.std(X, 0)
X_std = (X - mu) / sigma ## <-- SOLUTION
return X_std
X_std = standardise(X)
# check
X_std.tail()
| radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | symmetry_mean | fractal_dimension_mean | ... | radius_worst | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||||
| 926424 | 2.110995 | 0.721473 | 2.060786 | 2.343856 | 1.041842 | 0.219060 | 1.947285 | 2.320965 | -0.312589 | -0.931027 | ... | 1.901185 | 0.117700 | 1.752563 | 2.015301 | 0.378365 | -0.273318 | 0.664512 | 1.629151 | -1.360158 | -0.709091 |
| 926682 | 1.704854 | 2.085134 | 1.615931 | 1.723842 | 0.102458 | -0.017833 | 0.693043 | 1.263669 | -0.217664 | -1.058611 | ... | 1.536720 | 2.047399 | 1.421940 | 1.494959 | -0.691230 | -0.394820 | 0.236573 | 0.733827 | -0.531855 | -0.973978 |
| 926954 | 0.702284 | 2.045574 | 0.672676 | 0.577953 | -0.840484 | -0.038680 | 0.046588 | 0.105777 | -0.809117 | -0.895587 | ... | 0.561361 | 1.374854 | 0.579001 | 0.427906 | -0.809587 | 0.350735 | 0.326767 | 0.414069 | -1.104549 | -0.318409 |
| 927241 | 1.838341 | 2.336457 | 1.982524 | 1.735218 | 1.525767 | 3.272144 | 3.296944 | 2.658866 | 2.137194 | 1.043695 | ... | 1.961239 | 2.237926 | 2.303601 | 1.653171 | 1.430427 | 3.904848 | 3.197605 | 2.289985 | 1.919083 | 2.219635 |
| 92751 | -1.808401 | 1.221792 | -1.814389 | -1.347789 | -3.112085 | -1.150752 | -1.114873 | -1.261820 | -0.820070 | -0.561032 | ... | -1.410893 | 0.764190 | -1.432735 | -1.075813 | -1.859019 | -1.207552 | -1.305831 | -1.745063 | -0.048138 | -0.751207 |
5 rows × 30 columns
# split into train and test set
# stacking data X and labels y into one matrix
data_split = np.hstack((X_std, y[:, np.newaxis]))
# There can be a bug caused by Python versions, try also data_split = np.hstack((X_std, np.array(y)[:, np.newaxis]))
# shuffling the rows
np.random.shuffle(data_split)
# we split train to test as 70:30
split_rate = 0.7
train, test = np.split(data_split, [int(split_rate*(data_split.shape[0]))])
X_train = train[:,:-1]
y_train = train[:, -1]
X_test = test[:,:-1]
y_test = test[:, -1]
y_train = y_train.astype(float)
y_test = y_test.astype(float)
# insert 1 in every row for intercept b
X_train_intercept = np.hstack((X_train, np.ones((len(X_train),1)) ))
X_test_intercept = np.hstack((X_test, np.ones((len(X_test),1)) ))
/tmp/ipykernel_4930/2016496295.py:3: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
data_split = np.hstack((X_std, y[:, np.newaxis]))
Section 2: Linear SVM Formulation
We start with defining the hinge loss as
L ( w ) = 1 2 ∥ w ∥ 2 + λ ∑ i = 1 n max ( 0 , 1 − y i ( x ( i ) ⋅ w + b ) ) . \mathcal L (\boldsymbol w) = \frac{1}{2} \| \boldsymbol w \|^2 + \lambda \sum_{i=1}^n \max \bigg( 0, 1-y_i (x^{(i)} \cdot \boldsymbol w + b) \bigg) \, . L(w)=21∥w∥2+λi=1∑nmax(0,1−yi(x(i)⋅w+b)).
where w \boldsymbol w w is the vector of weights, λ \lambda λ the regularisation parameter, and b b b the intercept which is included in our X as an additional column of 1 1 1’s.
# EDIT THIS FUNCTION
def compute_cost(w, X, y, regul_strength=1e5):
"""
Arguments:
w: weight vector
X, y: data (features and target)
regul_strength: lambda
Returns:
loss: see equation above
"""
n = X.shape[0]
distances = 1 - y * (X @ w) ## <-- SOLUTION
distances[distances < 0] = 0 # equivalent to max(0, distance)
hinge = regul_strength * distances.mean() ## <-- SOLUTION
# calculate cost
return 0.5 * np.dot(w, w) + hinge - 0.5 * w[-1] ** 2
Section 3: SVM Optimization using SGD
One way to optimize the cost is by using stochastic gradient descent (SGD) algorithm. In order to use SGD, we need to implement a function for the cost gradients with respect to w \boldsymbol w w.
# calculate gradient of cost
def calculate_cost_gradient(w, X_batch, y_batch, regul_strength=1e6):
"""
Arguments:
w: weight vector
X_batch, y_batch: data (features and target) in a selected batch
regul_strength: lambda
Returns:
gradient of the loss w.r.t. w
"""
# if only one example is passed
if type(y_batch) == np.float64:
y_batch = np.asarray([y_batch])
X_batch = np.asarray([X_batch]) # gives multidimensional array
distance = 1 - (y_batch * (X_batch @ w))
dw = np.zeros(len(w))
we = w.copy() # So as not to overwrite w
we[-1] = 0 # So as not to have b in its derivative when adding the weights in di
for ind, d in enumerate(distance):
if max(0, d)==0: ## <-- SOLUTION
di = we ## <-- SOLUTION
else:
di = we - (regul_strength * y_batch[ind] * X_batch[ind]) ## <-- SOLUTION
dw += di
return dw/len(y_batch) # average
Both of the two previous functions are then used in SGD to update the weights iteratively with a given learning rate α \alpha α. We also implement a stop criterion that ends the learning as soon as the cost function has not changed more than a manually determined percentage.
We know that the learning happens through updating the weights according to
w = w − α ∂ L ∂ w \boldsymbol w = \boldsymbol w - \alpha \frac{\partial \mathcal L}{\partial \boldsymbol w} w=w−α∂w∂L
where ∂ L ∂ w \frac{\partial \mathcal L}{\partial \boldsymbol w} ∂w∂L is the gradient of the hinge loss we have computed in the previous cell.
# EDIT THIS FUNCTION
def sgd(X, y, max_iterations=2000, stop_criterion=0.01, learning_rate=1e-5, regul_strength=1e6, print_outcome=False):
"""
Arguments:
X, y: data (features and target)
max_iterations: maximum iteration of training
stop_criterion: stop if the percentage change in loss is less than this
regul_strength: lambda
print_outcome: whether to print the loss at certain iterations
Returns:
weights: w
cost_list: list of losses stored at certain iterations
iteration_list: list of iterations when you recorded the loss
"""
# initialise zero weights
weights = np.zeros(X.shape[1])
nth = 0
# initialise starting cost as infinity
prev_cost = np.inf
# record costs
cost_list = []
iteration_list = []
# stochastic gradient descent
indices = np.arange(len(y))
for iteration in range(max_iterations):
# shuffle to prevent repeating update cycles
np.random.shuffle(indices)
X, y = X[indices], y[indices]
for xi, yi in zip(X, y):
descent = calculate_cost_gradient(weights, xi, yi, regul_strength) ## <-- SOLUTION
weights = weights - (learning_rate * descent) ## <-- SOLUTION
# convergence check on 2^n'th iteration
if iteration==2**nth or iteration==max_iterations-1:
# compute cost
cost = compute_cost(weights, X, y, regul_strength) ## <-- SOLUTION
if print_outcome:
print("Iteration is: {}, Cost is: {}".format(iteration, cost))
# stop criterion
if abs(prev_cost - cost) < stop_criterion * prev_cost: ## <-- SOLUTION
return weights, cost_list, iteration_list
prev_cost = cost
iteration_list.append(iteration)
cost_list.append(cost)
nth += 1
return weights, cost_list, iteration_list
Now, we can take these functions and train a linear SVM with our training data.
# train the model
lam=10
w, costs, iterations = sgd(X_train_intercept, y_train, max_iterations=2000, stop_criterion=0.001, learning_rate=1e-5, regul_strength=lam, print_outcome=True)
print("Training finished.")
Iteration is: 1, Cost is: 4.839282882996163
Iteration is: 2, Cost is: 3.755514893535763
Iteration is: 4, Cost is: 2.8101315147850703
Iteration is: 8, Cost is: 2.1690723478315888
Iteration is: 16, Cost is: 1.756586593237897
Iteration is: 32, Cost is: 1.4932071331932155
Iteration is: 64, Cost is: 1.3640789504858404
Iteration is: 128, Cost is: 1.282672173658206
Iteration is: 256, Cost is: 1.2657878225118049
Iteration is: 512, Cost is: 1.2611293410604092
Iteration is: 1024, Cost is: 1.260458893322897
Training finished.
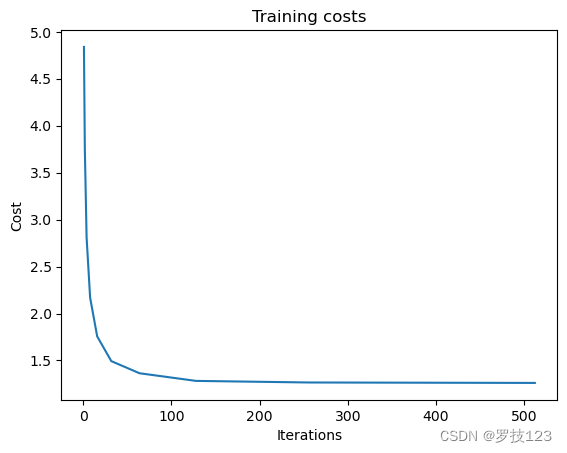
We can plot the cost against the iteractions to check nothing went wrong
plt.plot(iterations, costs)
plt.title('Training costs')
plt.xlabel("Iterations")
plt.ylabel("Cost")
plt.show()
To evaluate the mean accuracy in both train and test set, we write a small function called score.
## EDIT THIS FUNCTION
def score(w, X, y):
y_preds = np.sign(X @ w)
return np.mean(y_preds == y) ## <-- SOLUTION
print("Accuracy on training set: {}".format(score(w, X_train_intercept, y_train)))
print("Accuracy on test set: {}".format(score(w, X_test_intercept, y_test)))
Accuracy on training set: 0.9773869346733668
Accuracy on test set: 0.9649122807017544
Questions:
- What are other evaluation metrics besides the accuracy? Implement them and assess the performance of our classification algorithm with them.
- What makes other evaluation metrics more appropriate given our unbalanced data set (we have more benign than malignant examples)?
- Try different learning rates, regularisation strengths and number of iterations independently. What can you observe? Can you achieve higher accuracies?
- What is your understanding why have we used the hinge loss with this data set of 31 features?
Section 4: Model Evaluation via T-fold Cross Validation
Now we repeat the same procedure as above but do not only have one train-test split, but multiple in a T-fold cross validation method.
def cross_val_split(N, num_folds):
fold_size = N // num_folds
index_perm = np.random.permutation(np.arange(N))
folds = []
for k in range(num_folds):
folds.append(index_perm[k*fold_size:(k+1)*fold_size])
return folds
# evaluate
folds = cross_val_split(train.shape[0], 5)
folds
[array([ 84, 235, 170, 72, 64, 309, 108, 254, 222, 384, 325, 174, 106,
195, 246, 218, 221, 209, 194, 247, 273, 114, 26, 341, 355, 363,
145, 370, 66, 146, 157, 340, 160, 9, 304, 86, 267, 285, 385,
324, 358, 122, 308, 365, 76, 281, 396, 42, 109, 2, 100, 119,
289, 237, 149, 55, 88, 219, 376, 268, 291, 143, 18, 323, 98,
220, 391, 152, 231, 394, 163, 334, 353, 197, 364, 147, 296, 85,
190]),
array([196, 87, 356, 51, 20, 303, 395, 180, 189, 168, 45, 165, 213,
252, 47, 81, 269, 8, 199, 284, 388, 264, 204, 10, 299, 150,
92, 371, 171, 33, 104, 48, 28, 103, 320, 183, 121, 38, 127,
288, 297, 12, 377, 257, 344, 232, 73, 173, 23, 313, 31, 393,
138, 338, 310, 59, 74, 185, 131, 256, 166, 130, 345, 179, 0,
300, 101, 68, 137, 155, 133, 360, 240, 3, 58, 212, 302, 375,
362]),
array([367, 319, 280, 32, 390, 187, 77, 348, 111, 314, 214, 224, 359,
295, 113, 140, 233, 275, 102, 115, 191, 82, 116, 83, 330, 62,
144, 67, 215, 65, 162, 337, 120, 19, 4, 118, 139, 192, 333,
110, 244, 167, 381, 99, 293, 202, 153, 17, 315, 316, 154, 361,
40, 382, 369, 172, 6, 386, 148, 298, 354, 260, 94, 307, 368,
238, 343, 93, 346, 248, 317, 142, 136, 389, 331, 283, 241, 274,
125]),
array([169, 276, 272, 49, 203, 75, 287, 282, 251, 226, 255, 161, 216,
21, 63, 211, 95, 234, 328, 261, 11, 34, 335, 380, 53, 141,
378, 184, 134, 43, 27, 80, 1, 135, 60, 164, 156, 225, 112,
200, 107, 70, 262, 336, 239, 372, 91, 271, 54, 198, 259, 387,
286, 50, 207, 339, 294, 90, 205, 36, 61, 208, 392, 181, 253,
159, 193, 69, 349, 332, 236, 14, 22, 351, 311, 373, 96, 188,
201]),
array([186, 44, 243, 229, 89, 258, 210, 366, 245, 277, 305, 263, 306,
24, 158, 350, 329, 301, 16, 321, 39, 250, 56, 318, 178, 37,
79, 228, 397, 117, 7, 374, 326, 175, 29, 182, 15, 129, 177,
352, 52, 327, 13, 312, 123, 279, 290, 176, 25, 278, 342, 292,
206, 5, 265, 126, 46, 217, 322, 227, 223, 266, 249, 128, 151,
242, 383, 124, 35, 78, 270, 41, 357, 71, 57, 132, 230, 105,
347])]
## EDIT THIS FUNCTION
def cross_val_evaluate(data, num_folds):
"""
Arguments:
data: training data with which T-fold CV is performed
num_folds: T, the number of folds
Returns:
train_scores: train accuracy for each fold
val_scores: validation accuracy for each fold
"""
folds = cross_val_split(data.shape[0], num_folds)
train_scores = []
val_scores = []
for i in range(len(folds)):
print('Fold', i+1)
val_indices = folds[i]
# define the training set
train_indices = list(set(range(data.shape[0])) - set(val_indices))
X_train = data[train_indices, :-1] ## <-- SOLUTION
y_train = data[train_indices, -1]
# define the validation set
X_val = data[val_indices, :-1] ## <-- SOLUTION
y_val = data[val_indices, -1] ## <-- SOLUTION
# insert 1 in every row for intercept b
X_train = np.hstack((X_train, np.ones((len(X_train),1)) ))
X_val = np.hstack((X_val, np.ones((len(X_val),1)) ))
# train the model
w,_,_ = sgd(X_train, y_train, max_iterations=1024, stop_criterion=0.01, learning_rate=1e-5, regul_strength=1e3)
print("Training finished.")
# evaluate
train_score = score(w, X_train, y_train)
val_score = score(w, X_val, y_val)
print("Accuracy on training set #{}: {}".format(i+1, train_score))
print("Accuracy on validation set #{}: {}".format(i+1, val_score))
train_scores.append(train_score)
val_scores.append(val_score)
return train_scores, val_scores
train_scores, val_scores = cross_val_evaluate(train, 5)
Fold 1
Training finished.
Accuracy on training set #1: 0.9937304075235109
Accuracy on validation set #1: 0.9746835443037974
Fold 2
Training finished.
Accuracy on training set #2: 0.9843260188087775
Accuracy on validation set #2: 0.9873417721518988
Fold 3
Training finished.
Accuracy on training set #3: 0.9905956112852664
Accuracy on validation set #3: 0.9493670886075949
Fold 4
Training finished.
Accuracy on training set #4: 0.9905956112852664
Accuracy on validation set #4: 0.9620253164556962
Fold 5
Training finished.
Accuracy on training set #5: 0.987460815047022
Accuracy on validation set #5: 0.9873417721518988
Finally, let’s compute the mean accuracy.
print(np.mean(train_scores), np.mean(val_scores))
0.9893416927899686 0.9721518987341773
Subsection 4.1: Kernelised SVM
In the following, we implement a soft-margin kernelised SVM classifier with a non-linear kernel. Here, we use a Gaussian kernel:
k ( x , y ∣ σ ) = e − ∣ ∣ x − y ∣ ∣ 2 σ k(x,y|\sigma) = e^{-\frac{||x-y||^2}{\sigma}} k(x,y∣σ)=e−σ∣∣x−y∣∣2
# First we need a function to calculate the kernel given the data #
def kernel_matrix(X1,X2,sigma):
"""
Arguments:
X1, X2: two input datasets
sigma: hyperparameter in Gaussian kernel (look up for its interpretation if you are interested)
Returns:
kernel: K matrix
"""
n1,m1 = X1.shape
n2,m2 = X2.shape
kernel = np.zeros((n1,n2))
# Here we define a Gaussian Radial Basis Function Kernel #
for i in range(n1):
exponent = np.linalg.norm(X2 - X1[i],axis=1)**2 ## <-- SOLUTION
kernel[i,:] = np.exp(-exponent/sigma)
return kernel
Having defined the kernel, we use this to compute the cost below.
## EDIT THIS FUNCTION
def compute_cost_kernel(u, K, y, regul_strength=1e3,intercept=0):
"""
Arguments:
u, K, y: vector/matrix as defined in notes
regul_strength: lambda
intercept: b, to be estimated seperately
Returns:
loss: as defined below
"""
# Here I define the hinge cost with the kernel trick. NB: the intercept should be kept separate #
distances = 1 - y*(K@u + intercept) ## <-- SOLUTION
distances[distances < 0] = 0 # equivalent to max(0, distance)
hinge = regul_strength * distances.mean()
# calculate cost
return 0.5 * np.dot(u,K@u) + hinge # + 0.5 * intercept ** 2, if regularise b
As we have seen in the lecture notes, the kernel trick can be implemented in the primal-form problem by defining a new loss function:
L ( u , b ) = 1 2 u T K u + λ ∑ i = 1 N max { 0 , 1 − y ( i ) ( K ( i ) u + b ) } , L(\mathbf{u},b) = \frac{1}{2}\mathbf{u}^{\rm{T}}\mathbf{K} \mathbf{u} + \lambda \sum_{i=1}^N \max \Big\{0, 1-y^{(i)}(\mathbf{K}^{(i)}\mathbf{u} + b)\Big\}, L(u,b)=21uTKu+λi=1∑Nmax{0,1−y(i)(K(i)u+b)},
where K \mathbf{K} K is the matrix containing the kernel functions, i.e.
K i j = k ( x ( i ) , x ( i ) ) \mathbf{K}_{ij} = k(\mathbf{x}^{(i)},\mathbf{x}^{(i)}) Kij=k(x(i),x(i)). To perform the optimisation, we simply modify the functions introduced above. Note that we will use X t r a i n X_{\mathrm{train}} Xtrain and X t e s t X_{\mathrm{test}} Xtest without the additional vector of ones. We had previously included this vector of ones to learn the intercept term b b b, but within the new formulation, one cannot readily employ this trick. How to include the intercept in kernalised SVM was a question in CW1 2023, and this year we make it an exercise in this notebook.
Following the steps above, we want to optimize the cost is by using SGD. First, we thus create a function that computes the gradients.
## EDIT THIS FUNCTION
def calculate_cost_gradient_kernel(u, K_batch, y_batch, regul_strength=1e3,intercept=0):
"""
Arguments:
u, K_batch, y_batch: vector/matrix as defined in notes
regul_strength: lambda
intercept: b
Returns:
gradient of the loss w.r.t. u
"""
# if only one example is passed
if type(y_batch) == np.float64 or type(y_batch) == np.int32:
y_batch = np.asarray([y_batch])
K_batch = np.asarray([K_batch]) # gives multidimensional array
distance = 1 - (y_batch * (K_batch @ u + intercept)) ## <-- SOLUTION
dw = np.zeros(len(u))
db = 0
# define the gradient with the hinge loss #
for ind, d in enumerate(distance):
if max(0, d)==0:
di = K_batch@u ## <-- SOLUTION
di_intercept = 0 # di_intercept = intercept, if regularise b
else:
di = K_batch@u - (regul_strength * y_batch[ind] * K_batch[ind]) ## <-- SOLUTION
di_intercept = - regul_strength * y_batch[ind] ## <-- SOLUTION # + intercept, if regularise b
dw += di
db += di_intercept
return dw/len(y_batch), db/len(y_batch)
We can now use the functions above in SGD.
## EDIT THIS FUNCTION
def sgd_kernel(K, y, batch_size=32, max_iterations=4000, stop_criterion=0.001, learning_rate=1e-4, regul_strength=1e3, print_outcome=False):
"""
Arguments:
K, y: data (transformed features and target)
batch_size: size of a batch in SGD
max_iterations: maximum iteration of training
stop_criterion: stop if the percentage change in loss is less than this
regul_strength: lambda
print_outcome: whether to print the loss at certain iterations
Returns:
weights: u
intercept: b
"""
# initialise zero u and intercept
u = np.zeros(K.shape[1])
intercept=0
nth = 0
# initialise starting cost as infinity
prev_cost = np.inf
# stochastic gradient descent
indices = np.arange(len(y))
for iteration in range(max_iterations):
# shuffle to prevent repeating update cycles
np.random.shuffle(indices)
batch_idx = indices[:batch_size]
K_b, y_b = K[batch_idx], y[batch_idx]
for ki, yi in zip(K_b, y_b):
ascent, ascent_intercept = calculate_cost_gradient_kernel(u, ki, yi, regul_strength, intercept) # <-- SOLUTION
u = u - (learning_rate * ascent)
intercept = intercept - (learning_rate * ascent_intercept) ## <-- SOLUTION
# convergence check on 2^n'th iteration
if iteration==2**nth or iteration==max_iterations-1:
# compute cost
cost = compute_cost_kernel(u, K, y, regul_strength, intercept) ## <-- SOLUTION
if print_outcome:
print("Iteration is: {}, Cost is: {}".format(iteration, cost))
# stop criterion
if abs(prev_cost - cost) < stop_criterion * prev_cost: ## <-- SOLUTION
return u, intercept
prev_cost = cost
nth += 1
return u, intercept
Finally, let’s compare some differnet values of σ.
## EDIT THIS FUNCTION
for sigma in [1,2,5,10]:
print('For sigma = ' + str(sigma))
K_train = kernel_matrix(X_train,X_train, sigma) # note X_train doesn't have 1's attached
u,b = sgd_kernel(K_train, y_train, batch_size=128, max_iterations=2000, stop_criterion=0.001, learning_rate=1e-5, regul_strength=1e3, print_outcome=False)
def score(u, X, y, sigma, intercept):
## now I define the kernel containing test and train data ##
K_test = kernel_matrix(X, X_train, sigma)
## The
y_preds = np.sign(K_test@u + intercept) ## <-- SOLUTION
return np.mean(y_preds == y)
print("Accuracy on training set: {}".format(score(u, X_train, y_train, sigma, b)))
print("Accuracy on test set: {}".format(score(u, X_test, y_test, sigma, b)))
print("Intercept: {}".format(b))
For sigma = 1
Accuracy on training set: 1.0
Accuracy on test set: 0.6608187134502924
Intercept: -0.19999999999999996
For sigma = 2
Accuracy on training set: 1.0
Accuracy on test set: 0.7251461988304093
Intercept: -0.27
For sigma = 5
Accuracy on training set: 1.0
Accuracy on test set: 0.8362573099415205
Intercept: -0.42000000000000026
For sigma = 10
Accuracy on training set: 0.9974874371859297
Accuracy on test set: 0.9473684210526315
Intercept: -0.14000000000000007
Questions:
- What do you observe varying sigma?
- How does the Gaussian Kernel SVM compare to the linear SVM?
Section 5: How to make a nice plot …
In your coursework, we ask you to produce some plots. Here, we outline how a “good” plot looks. For this purpose, let’s consider a specific example based on the data in this notebook. Thus, let’s plot the diagnosis (malignant or benign) in a scatter plot as a function of, say, the two first features (radius_mean and texture_mean). If we just plot the data using the standard settings, we don’t end up with a very good plot.
plt.scatter(data_2.iloc[:, 1],data_2.iloc[:, 2],c=y);
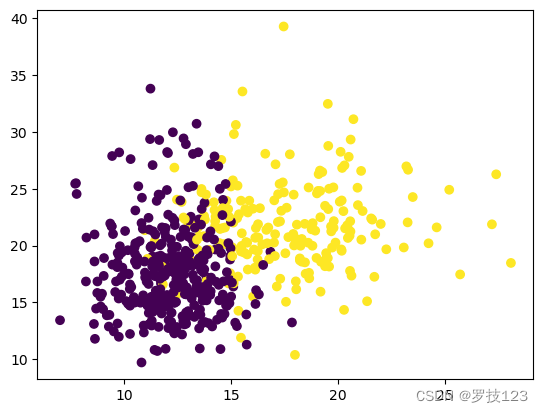
The problem is the following: As a rule of thumb, a plot should always be able to stand alone. In other words, it should be possible to understand the plot without reading the surrounding code or text and even (if possible) without reading the figure caption. The plot above can clearly not stand on its own. For starters, it doesn’t have any legend, nor does it have labels on the axes. Likewise, we should add a title. Other types of plots might have other options to add information. For instance, if we were dealing with a contour plot, we might want to include a colour bar or add labels to the contour lines.
labels = ["Malignant", "Benign"]
for i, c in enumerate(np.unique(y)):
plt.scatter(data_2.iloc[:, 1][y==c],data_2.iloc[:, 2][y==c],label=labels[i]);
plt.xlabel("Mean Radius");
plt.ylabel("Mean Texture");
plt.title("Clustering of Diagnoses");
plt.legend();
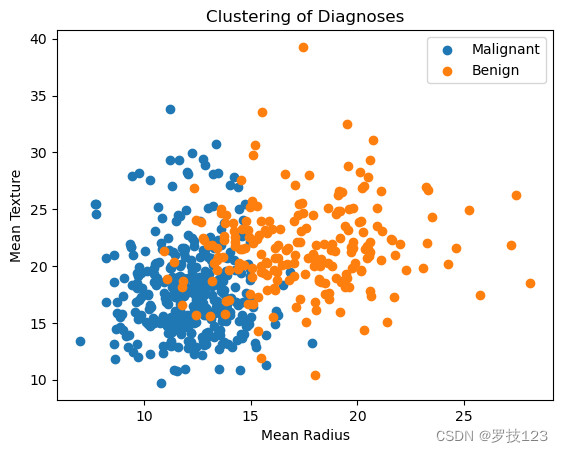
These small additions have improved the plot significantly. But it’s still not optimal. First of all, it’s rather small. To make it more readable, we should increase the figure size and change the font size of the different labels, titles, and ticks. Now, what is a suitable font size? As a rule of thumb, a plot should be easy to read when you print it. If you printed the notebook, the plot would be a bit on the small side since the text is smaller than the text in the cells; thus, if someone scrolls through your notebook, they would probably have to zoom in.
Secondly, we might want to reconsider the markers. Indeed, there is an addition to the rule stated above: A plot should be easy to read when you print it in grayscale. The idea is that you make the plot colourblind-friendly if you ensure that your plot is grayscale convertible. Thus, use colormaps that have this property. The default colormap of plt.scatter is actually grayscale convertible, leading to the choice of yellow and purple markers in the first plot in this section. While the notebook changed colours when creating the second plot, it chose colours that were “luckily” likewise grayscale convertible. If in doubt, you can, of course, use your own colour scheme, as shown below. Another way to make the plot more readable when printed in grayscale (and hence more readable in general) is to use different markers in addition to colours that are easily distinguishable. We, therefore, also change the markers themselves below. You can use the same trick when you make other plots. If you were to make, say, a line plot, you could consider, e.g., using different line styles (“-”,“:”,“–”).
fig, ax = plt.subplots(figsize=(12,8)) # We make plot larger
markers = ["^","o"] # We use distinct markers
labels = ["Benign", "Malignant"]
col = ['#fde724','#440154']; # We want to change back to the yellow and purple used above.
fsize = 14 # We set fontsize
for i, c in enumerate(np.unique(y)):
ax.scatter(data_2.iloc[:, 1][y==c],data_2.iloc[:, 2][y==c],c=col[i],marker=markers[i],label=labels[i],s=50);
ax.legend(loc="best",fontsize=fsize);
plt.xlabel("Mean Radius",fontsize=fsize+4);
plt.ylabel("Mean Texture",fontsize=fsize+4);
plt.xticks(size=fsize); # We change the size of the ticks
plt.yticks(size=fsize); # We change the size of the ticks
plt.title("Clustering of Diagnoses",fontsize=2*fsize);
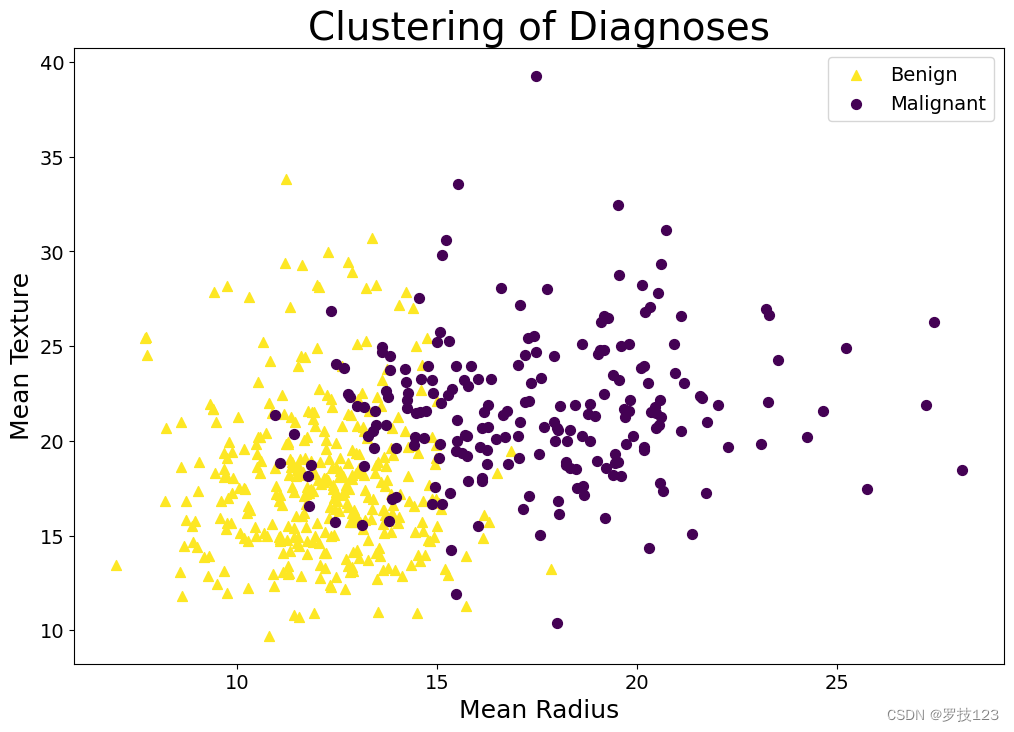
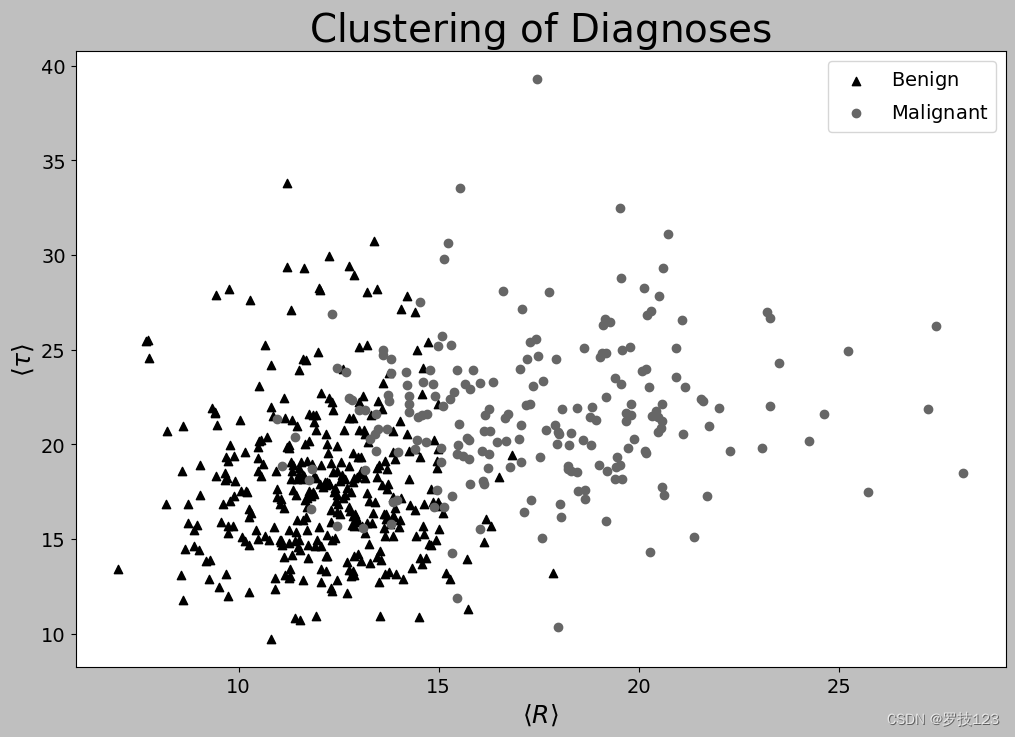
The plot above is what we would consider a “good” plot since it fulfils both the criteria sketched above: It can stand alone, and it’s easy to read. But what if we had decided to plot something different, such as the regularisation for the kernelised SVM, i.e. sigma? In that case, LaTeX commands would be useful for writing σ \sigma σ rather than “sigma”. Indeed, we can readily do so in Python by passing a raw string (r"") with a LaTeX command. So, for the sake of the example, let’s say that we have called the radius R and the texture τ \tau τ.
# The plots in your report should be in colour. But just to show what we mean with grayscale convertable...
plt.style.use('grayscale')
fig, ax = plt.subplots(figsize=(12,8))
markers = ["^","o"]
labels = [r"$\mathrm{Benign}$", r"$\mathrm{Malignant}$"] # We now use LaTeX commands
fsize = 14 # Change fontsize
for i, c in enumerate(np.unique(y)):
ax.scatter(data_2.iloc[:, 1][y==c],data_2.iloc[:, 2][y==c],marker=markers[i],label=labels[i]);
ax.legend(loc="best",fontsize=fsize);
plt.xlabel(r"$\langle R \rangle$",fontsize=fsize+4);
plt.ylabel(r"$\langle \tau \rangle$",fontsize=fsize+4);
plt.xticks(size=fsize);
plt.yticks(size=fsize);
plt.title(r"$\mathrm{Clustering \,\, of \,\, Diagnoses}$",fontsize=2*fsize);



















![树形DP——[CTSC1997]选课](https://img-blog.csdnimg.cn/direct/13e95c95aed9488fa2db2a2ca59add8e.png)