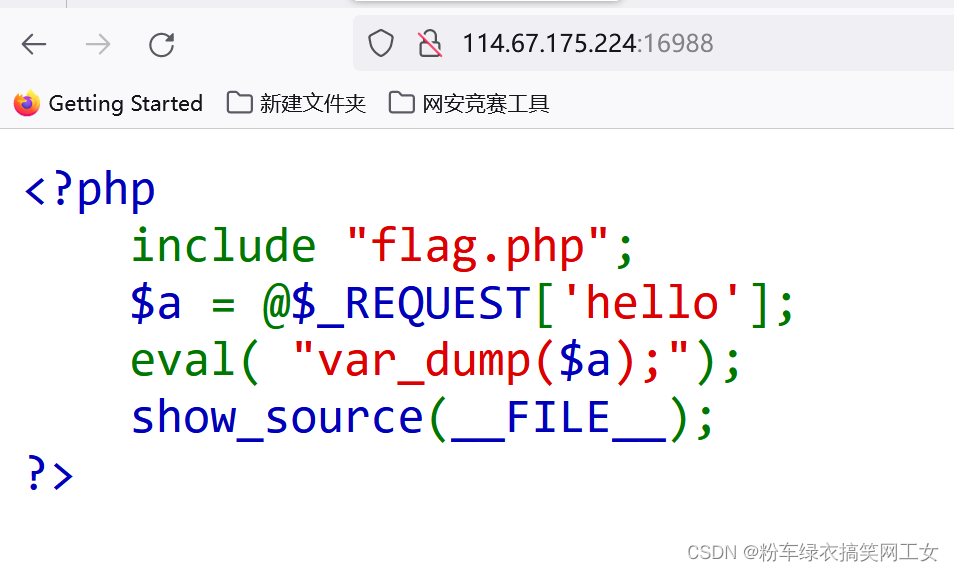
1.假如下面的shell脚本"2024_extractrG4_calculating_score_filternonoverlapG4.sh"
#!/bin/bash
python lin_change_gff3original.py -i iwgsc_refseqv2.1_annotation_200916_HC_LC.gff3 -o iwgsc_refseqv2.1_annotation_200916_HC_LC.gff4
# Define base filenames
fasta_base="iwgsc_refseqv2.1_annotation_200916_HC_LC_mrna.fasta"
gff_file="iwgsc_refseqv2.1_annotation_200916_HC_LC.gff4"
# Array of types
types=("G4" "C4" "A4" "T4")
# Loop through each type and process files accordingly
for type in "${types[@]}"
do
# Python script to extract r[type] structures
python "lin_20240425_extract_r${type}.py" -f1 "$fasta_base" -f2 "lijinonextended_mrna_allr${type}output.fasta"
# R script to calculate r[type] score
Rscript "lin_20240321_calculating_r${type}score.R" -i "lijinonextended_mrna_allr${type}output.fasta" -o "lijinonextended_mrna_allr${type}output1.fasta"
# Python script to annotate with gff4
python "lin_annotating_r${type}_with_gff4.py" -g "$gff_file" -f "lijinonextended_mrna_allr${type}output1.fasta" -o "lijinonextended_mrna_allr${type}output2.fasta"
less "lijinonextended_mrna_allr${type}output2.fasta" | grep -E 'Id|five' > "five_r${type}_total.txt"
less "lijinonextended_mrna_allr${type}output2.fasta" | grep -E 'Id|CDS' > "cds_r${type}_total.txt"
less "lijinonextended_mrna_allr${type}output2.fasta" | grep -E 'Id|three' > "three_r${type}_total.txt"
cp "lijinonextended_mrna_allr${type}output2.fasta" "mrna_r${type}_total.txt"
python lin_filter_non-overlap_rG4.py -f1 "five_r${type}_total.txt" -f2 "five_r${type}_total1.txt"
python lin_filter_non-overlap_rG4.py -f1 "cds_r${type}_total.txt" -f2 "cds_r${type}_total1.txt"
python lin_filter_non-overlap_rG4.py -f1 "three_r${type}_total.txt" -f2 "three_r${type}_total1.txt"
python lin_filter_non-overlap_rG4.py -f1 "mrna_r${type}_total.txt" -f2 "mrna_r${type}_total1.txt"
done
如果,我后台运行这个脚本之后,想取消总的进程,可以这样操作:
ps aux | grep 2024_extractrG4_calculating_score_filternonoverlapG4.sh
kill -SIGINT 2852911